ReSurfX::vysen Yields Remarkably Accurate and Actionable Insights Using System Response to Triggers – A Drug Response Study
Recently we added a new ‘System Response Based Triggers and Outcomes Predictor’ (SyRTOP) to our enterprise grade Decision Analytics software product ReSurfX::vysen. Here we share an exemplary result that proves the enhanced accuracy, lack of contaminating incorrect results, and many other advantages of this new solution. SyRTOP was tested on a system-wide gene expression response in liver tissue after treatment with a panel of drugs compared against a larger drug response database (Knowledge Repository) and proved to be a best in class solution as with other solutions incorporated in ReSurfX::vysen. The result is shown below.
click on the image to see at high resolution
Update: February 13, 2021: A newer article on ReSurfX::vysen in SyRTOP configuration with further improved results, powerful validations and description of components
ReSurfX in 2021 – Best-in-class Outcome Predictors, Innovation Catalysts and ROI Multipliers
is published on February 11, 2021. This article has some information complementary to that.
We have been taking the approach of building turnkey products and testing them in large-scale applications (de-risking the product and technology at scale) in comparison to widely used public domain and commercial solutions. We have also been learning from customer use of our products in a variety of ongoing data-intensive analytics initiatives.
We continue to release information in the ongoing Whitepaper Series ‘Improving Outcomes Through Enhanced Data Analytics and Artificial Intelligence, where we highlight the superior accuracy and the unique insights facilitated by our novel machine learning solution Adaptive Hypersurface Technology (AHT) that cannot be easily achieved using other machine learning techniques. The whitepaper series also showcase the capabilities of AHT-enabled solutions in our product ReSurfX::vysen1 for automated knowledge extraction. The release of results like the one here or in our article ‘Overcoming the Curse of Dimensionality with Combinatorics’ are driven by their timeliness and importance. As with the previous solutions incorporated into ReSurfX::vysen, SyRTOP proved to be a best in class solution that we are showcasing to our current and potential customers and stakeholders. Given that the novel machine learning approach that we invented - Adaptive Hypersurface Technology ‘AHT’ - is data-source agnostic, the example shown below relating to predicting drug mechanism of action or treatment outcomes has broad implications and uses with a variety of response and outcome prediction scenarios.
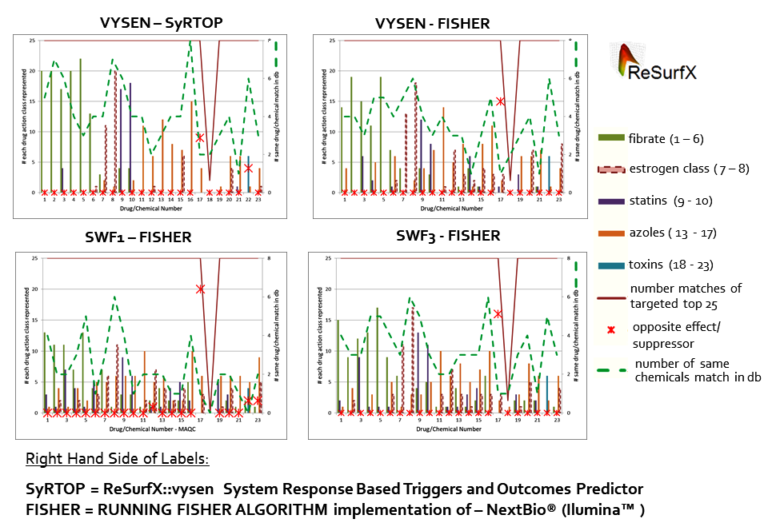
Let us dive directly into the most recent results shown above. Each of the four graph modules in the figure show the ability to predict mode of action by SyRTOP based on drug classes(es) of the closest matches in the target database (Knowledge Repository) of system-wide gene expression data to a query ‘system response’. In this example, system-wide gene expression data from toxicogenomic study data generated by the Microarray Quality Control Consortium for 23 compounds were used as query and were compared to a larger set of 678 gene expression response profiles to drug treatments. The 678 profiles used here consists of responses analyzed from data measured using microarray and sequencing technologies in liver tissues (i) generated by the Microarray and Sequencing quality control consortia (MAQC & SEQC)2, and (ii) 650 additional responses generated by the US National Toxicological Program (NTP)3 using 67 commercial drugs and other relevant chemicals at different concentrations and durations of exposure – in each case compared against relevant control liver tissues.
The best result is the one shown on the top left panel (labelled VYSEN – SyRTOP), far superior to the other three panels). This panel shows superior accuracy and cleanliness in determining drug classes (i.e., the known use of the system response being queried to the major known use of the closest matches). That result was generated using our new ‘System Response Based Triggers and Outcomes Predictor’ (SyRTOP, that we also refer to here as a COMPARATOR) to predict mechanism of action of drugs and outcomes of treatments. In this panel, both the system-wide gene response to the drug treatments used as inputs to SyRTOP queries as well as the database (Knowledge Repository) of gene expression responses to drug treatments were both generated by the AHT enabled solution in ReSurfX::vysen product.
The other three panels in the figure use another powerful COMPARATOR, the Running Fisher algorithm4, used in the commercial product NextBio®, now part of the sequencing instrumentation company Illumina™. In these latter three panels, differential gene expression was determined using either the ReSurfX::vysen product (top right panel) or two other approaches (bottom two panels). The other two approaches are (i) a widely used public domain solution and (ii) a commercial solution that are fed as input information to the Running-Fisher COMPARATOR implementation of NextBio® (both for the queried response and in the backend knowledge repository).
The results shown in each of the four panels in the figure can be summarized as two broad steps and the four graphical modules are labelled using a ‘Step 1 – Step2’ convention (e.g., VYSEN – VYSEN, VYSEN – FISHER etc.):
- Step 1: Identification of genes whose expression levels (collected by microarrays and sequencing) are affected by the drug treatments in liver tissue compared to relevant controls. These are data from a small number of replicates, which is a problem for most analytical approaches. Here we used either the ReSurfX::vysen product4 (labelled VYSEN), an implementation of a widely used public domain solution (SWF1)5 that includes the use of empirical Bayes, modeling, false discovery reduction, a variety of filters etc., or (SWF3)4 a published approach used in a commercial solution from NextBio® - now part of Illumina™. More detailed information at ReSurfX vysdom blog and whitepaper series.
- Step 2: The outputs from Step 1 are fed as input into the COMPARATORs, either ReSurfX SyRTOP or the Fisher algorithm3 that is incorporated into the NextBio® package. These COMPARATORs determine the proximity of the system response when compared to all 678 system responses to drugs.
Rather than using the data to make a predictor subset of parameters for each drug class, we opted to compare each gene expression response profile after the first step to the response profiles of all of the other treatments in the Knowledge Repository (backend database of system of response to treatments).
As mentioned above, the panel in the Figure labelled VYSEN-SyRTOP on the top left is absolutely clean in terms of the drug class(es) represented by the closest matches (top 25 matches when possible, or fewer as depicted by line graphs in brown) to the system responses being evaluated. The next best outcome is when comparative analytics data (Step 1) generated by ReSurfX::vysen was fed to the Running Fisher Algorithm COMPARATOR of NextBio®(Step 2) (labelled VYSEN-FISHER) (top right panel). This exemplifies the high quality of the input data provided by ReSurfX::vysen can improve results even from substandard steps of a workflow.
Thus the ReSurfX::vysen solution provides significant (non-incremental) advantages in two ways: (i) how data is analyzed to be used as input into a knowledge gleaning workflow, as well as (ii) in gleaning relevant knowledge from data. The other two input workflows when used in conjunction with the version of Running-Fisher-Algorithm (used in NextBio®/Illumina™ products) were suboptimal or in most cases picked the dominant response (primary mode of action of the drug being queried) a fewer number of times than the sum of the other signals (meaning, other drug classes). In other words, all other combinations of Step 1 and Step 2 have a lot more noise and incorrect signals in comparison to the VYSEN-SyRTOP analysis. The expression difference determination analytics (Step1: SWF1 and SWF3) and the response comparison scheme (Step 2: Running Fisher approach of NextBio®) used here to compare with solutions of ReSurfX::vysen are among the most widely used approaches, and are better than numerous other alternatives we have tested.
During the course of product development, we also found that that the current backend databases used for gleaning insights contain over 30% errors. Importantly: we have developed Big Data solutions to generate accurate custom Knowledge Repositories for large scale data sources of enterprise customers to overcome this high an error rate.
The drug response comparison use case shown here also has broader utility because it predicts responses that are most similar to each other from the whole system response (i.e., as an outcome of known and unknown effects and their interactions). We used the knowledge of the drug classes to show the power and accuracy of the solutions we are building into our products. The major classes of drugs in this subset are labelled in the figure and listed below:
- Fibrates: control/reduce ‘bad’ lipids that reduce cholesterol and reduce risk of heart disease and stroke and are used widely in people with Type 2 diabetes. The major mode of action of Fibrates is through activation of PPAR signaling pathway.
- Statins: another class of lipid lowering drug that reduce illness and mortality in those at risk of heart attack and stroke Statins mainly act through inhibiting HMGCoA Reductase.
- Estradiols: are used in birth control medications and to treat symptoms associated with menopause, prostrate and breast cancer, and to prevent osteoporosis. These medications act by affecting estrogen levels.
- Azoles: are primarily used as anti-fungal medications and to treat yeast infections. They act through P450 enzymes to inhibit ergosterol, a component of the cell membrane of target fungi and yeasts. Of the anti-fungal azoles represented here, only fluconazole (compound #15) is a triazole.
- Biological and Chemical Toxins: These include aflatoxins and beta-naphthoflavones, which have various forms of toxicities. They act in a number of known and unknown ways including DNA damage, Ah Receptor Agonists, and free radical generation.
- Two others: The 23 drugs also contained one azole-class drug used to treat hyperthyroidism and an anticonvulsant. A known pathway affected by these drugs is the CAR/PXR xenobiotic sensing system. Compounds #11 and #12 in the figure.
It is a well-known fact that all drugs have side effects. Even in the small subset of drugs used to highlight the power of SyrTOP in this article:
- statins and fibrates address common (but not the same) and overlapping drug use indications,
- anti-fungal azoles have estrogen modulating activity - the knowledge of which led to the development of optimized azole compounds to treat a number of estrogen dependent breast cancers (e.g., the widely used breast cancer drugs letrozole and anastrazole, both belong to triazole class).
ReSurfX::vysen SyRTOP seems to have an inherent ability to identify closer overall functionality of the intended use of the drug and has more control against favoring simple abundance of common features as the response of a drug. We make this inference from the system responses in the Knowledge Repository, that ranked high overall were not just same drug classes, but rather those variations of chemicals (drugs) that are optimized for one of its indicated treatment effects get picked first. A simple example is the fact that SyRTOP ranks matches of direct and optimized estrogen modulator drugs in the knowledge repository above those of azoles, even though azoles also modulate estrogens, when queried with the system response after treatment with an estrogen modulator. Observation of finer aspects (e.g., common side effects) of the drug action knowledge in the matches found also confirms this inference.
We made broad observations from over 42 million drug-drug treatment comparisons (each drug with its own set of biological replicates, and often matched, control group) using the drug response knowledge repository (KR) generated and contained in the ReSurfX::vysen product. We found that the VYSEN-SyRTOP combination help glean many useful insights well supported by medical knowledge and research literature, as well as new insights. The implications of the observations include:
- Given such a level of specificity, the new information obtained has a lot more significance (novel insights) and it is worth spending extra effort and resources (and how time is used) on ReSurfX::vysen-generated findings.
- More interestingly, the example of predicting drug classes and action from system-wide response in this study highlights the power of ReSurfX::vysen, even from one data source (gene expression measured using different technologies).
- In contrast to nearly all other solutions available, the ability of ReSurfX::vysen to be able to accomplish this degree of accuracy and predictive ability is very significant, especially from data with low levels replication (a common reason stated as failure of other statistical and machine learning approaches).
The data-source agnostic AHT leveraged solutions in our products for many different applications are equally powerful in combining many different types of data sources, as they are when only using gene expression data (from two measurement platforms) in the use case shown above.
Broader Needs that ReSurfX, ReSurfX::vysen and Adaptive Hypersurface Technology (AHT) Solve and Some Features of the Enterprise Software
The ReSurfX::vysen database subset used here is part of a larger Knowledge Repository contained in the ReSurfX::vysen product. During this process we have developed Big Data tools-based capability to generate custom large scale accurate Knowledge Repositories for customers from their proprietary data. As with the initial gene expression analytics leveraging our novel machine learning approach, the data-source agnostic Adaptive Hypersurface Technology (AHT) incorporated in ReSurfX::vysen, the new ‘System Response Based Triggers and Outcomes Predictor’ (SyRTOP) that enabled the above result, maintains our trend of building ‘best in class products’. The product also has a variety of features for enterprise applications, including:
- APIs and SDKs for seamless integration with customer workflows.
- In-built features for enhanced regulatory compliance and enterprise knowledge continuity.
- Incorporating powerful tools from other top technology companies to enable fast and accurate transfer of large amount of information.
- Option to integrate proprietary prediction approaches (COMPARATORs) of our customers as software plug-in to ReSurfX::vysen.
We have so far provided compelling evidence for the use of the capabilities of AHT and ReSurfX::vysen to significantly improve accuracy, provide novel insights, build better knowledge repositories with which to make better decisions, and help automate knowledge extraction thus improving overall innovation, outcomes and ROI.
We leverage some of the advantages highlighted here as well as other unique advantages of data-source agnostic Adaptive Hypersurface Technology (AHT) in our development of products for discovery through product sale solutions for Pharma/Biotech enterprises and novel solutions for predicting patient outcomes and evidence-based medicine for healthcare enterprises. Both these classes of applications support value-based business models and effective delivery of high cost treatments.
More information about ReSurfX offerings is available at https://resurfx.com and a rapid introduction at https://resurfx.com/learn-quick.
References:
- https://resurfx.com
- https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE47792
- https://ntp.niehs.nih.gov/data/drugmatrix/
- https://doi.org/10.1093/nar/gkv007
You can sign up to use the ReSurfX::vysen product at https://resurfx.com/vysen-signup/ and reach the ReSurfX team to utilize our solutions and products in your setting at https://resurfx.com.
Special thanks: Suresh Gopalan, Fred Ausubel, John Janakiraman, Saranya Balaji.
CHECK OUT OUR OTHER BLOGS AND WHITE PAPERS HERE.
ReSurfX in 2021 - Best-in-class Outcomes predictors, Innovation Catalysts and ROI Multipliers | ReSurfX, Feb 14, 2021
[…] result shown in this version is further improved than indicated in the introduction of SyRTOPReSurfX::vysen Yields Remarkably Accurate and Actionable Insights Using System Response to Triggers …dated Januaru 21, 2020 that already had marked improvement over other […]